TWAS coupled with eQTL analysis reveals the genetic connection between gene expression and flowering time in Arabidopsis
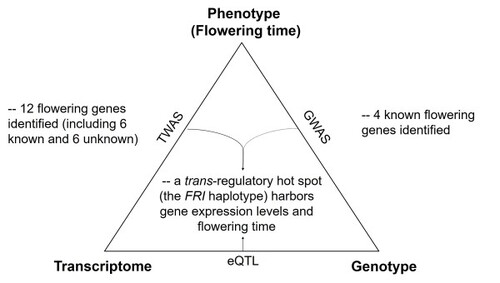
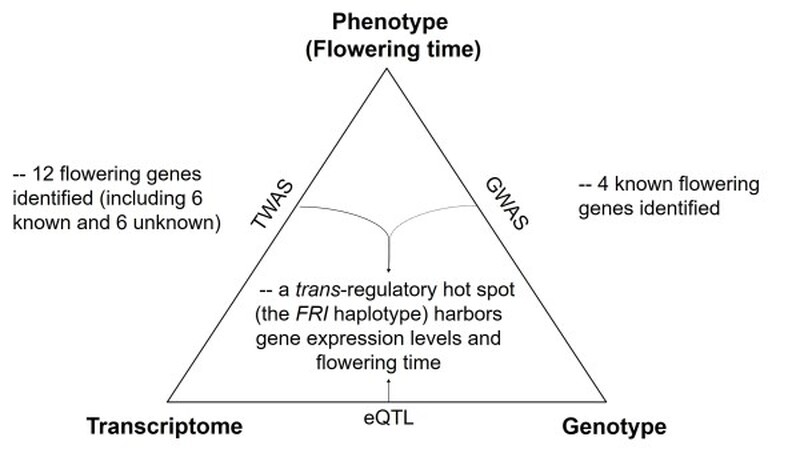
Genome-wide association study (GWAS) has improved our understanding of complex traits, but challenges exist in distinguishing causation versus association caused by linkage disequilibrium. Instead, transcriptome-wide association studies (TWAS) detect direct associations between expression levels and phenotypic variations, providing an opportunity to better prioritize candidate genes. To assess the feasibility of TWAS, we investigated the association between transcriptomes, genomes, and various traits in Arabidopsis, including flowering time. The associated genes formerly known to regulate growth allometry or metabolite production were first identified by TWAS. Next, for flowering time, six TWAS-newly identified genes were functionally validated. Analysis of the expression quantitative trait locus (eQTL) further revealed a trans-regulatory hotspot affecting the expression of several TWAS-identified genes. The hotspot covers the FRIGIDA (FRI) gene body, which possesses multiple haplotypes differentially affecting the expression of downstream genes, such as FLOWERING LOCUS C (FLC) and SUPPRESSOR OF OVEREXPRESSION OF CO 1 (SOC1). We also revealed multiple independent paths towards the loss of function of FRI in natural accessions. Altogether, this study demonstrates the potential of combining TWAS with eQTL analysis to identify important regulatory modules of FRI-FLC-SOC1 for quantitative traits in natural populations.
Pei-Shan Chien, Pin-Hua Chen, Cheng-Ruei Lee and Tzyy-Jen Chiou (2023) TWAS coupled with eQTL analysis reveals the genetic connection between gene expression and flowering time in Arabidopsis Journal of Experimental Botany, erad262, https://doi.org/10.1093/jxb/erad262

